Contact
Biography
I obtained my undergraduate degree in veterinary medicine from the University of La Plata, Argentina in 1996 and earned a PhD (Suma Cum Laude) from the University of Munich, Germany in 2001. My first postdoctoral training was done under supervision of Prof. Keith Campbell (2001-2003). In 2002 I was awarded a Marie Curie Fellowship (2002-2004) followed by an RCUK Fellowship (2005-2010). In 2010 I took up a position as Lecturer and was later promoted to Associate Professor (2015) and to Professor of Developmental Biology (2019) at the Division of Animal Sciences, School of Biosciences, UoN.
Expertise Summary
Our laboratory studies how cells become specialized during the formation of the fetus. We have special interest in the development of the precursors of the gametes, sperm and egg. We have used the pig embryo as a model system for understanding processes shared with human embryos. We have identified a key molecular program, based on the activation of the gene SOX17, in the precursors of the germ line. We have also identified sets of genes that escape DNA demethylation that are shared with those found in the human germline. These loci have the potential to carry epigenetic information across generations. The mechanisms underlying these processes are of considerable interest for understanding the basis of transgenerational inheritance.
For gaining detailed understanding of these events our laboratory uses molecular techniques that include single cell transcriptomics and bioinformatics. We also perform genome wide epigenetic analysis using PBAT and ATAC seq. We combine high-throughput technologies with gene editing using CRISPR/Cas9 systems to perform functional investigation in in vitro differentiation models as well as embryo editing.
We also have interest in understanding the molecular regulation of stemness in domestic animal embryos. We have used single cell sequencing techniques to define the molecular program of early pig embryos ans used this knowledge to derive embryonic stem cells and propagate them long term. These cells can give rise to all cell lineages. We have focussed in the derivation of muscle and fat progenitors with the aim of generating muscle in vitro. The purpose of this platform is to enhance the differentiation of these cells towards lab-grown meat, but also contribute to generating models of muscle for functional analysis of specific gene loci with high breeding value.
Teaching Summary
Biotechnology in Animal Physiology (BIOS3110; Co-Module convenor)
Animal Biotechnology (BIOS4122)
Regulation and Organisation in Animals (BIOS2088)
M.Med Sci in Assisted Reproductive Technology (A117)
Research Summary
Our laboratory investigates the developmental mechanisms of early mammalian embryos. Our aim is to gain understanding of the molecular regulation of cell decisions between the period when the… read more
Selected Publications
RAMOS-IBEAS P, SANG F, ZHU Q, TANG WWC, WITHEY S, KLISCH D, WOOD L, LOOSE M, SURANI MA and ALBERIO R, 2019. Pluripotency and X chromosome dynamics revealed in pig pre-gastrulating embryos by single cell analysis. Nature communications. 10(1), 500 KOBAYASHI T, ZHANG H, TANG WWC, IRIE N, WITHEY S, KLISCH D, SYBIRNA A, DIETMANN S, CONTRERAS DA, WEBB R, ALLEGRUCCI C, ALBERIO R and SURANI MA, 2017. Principles of early human development and germ cell program from conserved model systems. Nature. 546(7658), doi:10.1038/nature22812 JOHNSON AD and ALBERIO R, 2015. Primordial germ cells: the first cell lineage or the last cells standing? Development (Cambridge, England). 142(16), 2730-9 VALDEZ MAGAÑA G, RODRÍGUEZ A, ZHANG H, WEBB R and ALBERIO R, 2014. Paracrine effects of embryo-derived FGF4 and BMP4 during pig trophoblast elongation. Developmental biology. 387(1), 15-27
Current Research
Our laboratory investigates the developmental mechanisms of early mammalian embryos. Our aim is to gain understanding of the molecular regulation of cell decisions between the period when the blastocyst forms and the onset of gastrulation. This is a key period when the key lineages that make the adult organism are first set in the fetus and the placenta forms. Understanding how these processes are controlled in vivo can help us improve embryo survival and develop better technologies for the generation of useful cell types in vitro with uses in regenerative medicine and livestock production.
Our lab is embedded within CeLAB, a unique UK resource dedicated to research in Large Animal Biotechnology.
The current main research areas in our lab include:
Germ cell development in mammals
Germ cells are responsible for passing the genetic information of the parents to the next generation. The developmental program of these cells involves unique mechanisms that erase epigenetic memory (or epimutations) that may otherwise be passed on to the next generation. Our laboratory has shown that the germ cells in human and pig embryos are specified via similar mechanisms, thus making investigations in pig embryos of relevance to understanding human development (Kobayashi et al., Nature 416: 426-30, 2017; Zhu et al., Cell Reports 34:108735, 2021). Large mammals (humans, pigs, sheep and cows) have slower development, suggesting that the mechanisms for the establishment of different cell lineages, including the germline, may be under different control than those of mice. An important difference between the embryos of mice and other mammalian species, is that mouse embryos develop as a conical structure, which implies that cellular movements are very characteristic for this type of embryo. Other mammals develop as a flat embryonic disc, which means that the cellular movements are completely different to rodents.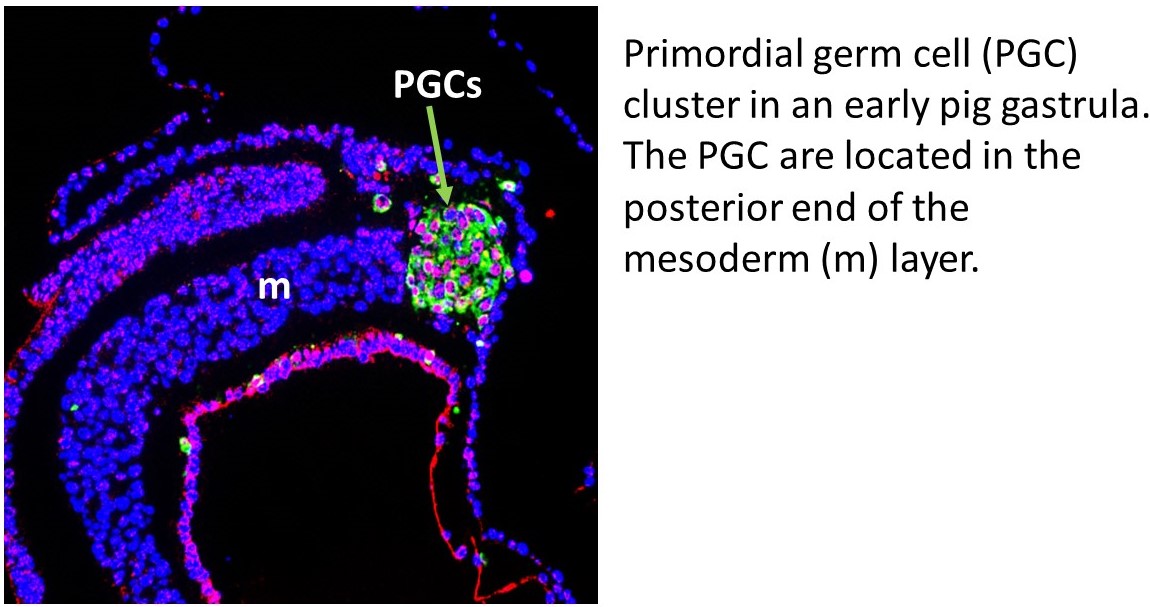 Given that many aspects of early pig embryo development are similar to human embryos, we use these embryos to uncover fundamental mechanisms of germ line development with relevance to human. The findings from our in vivo characterization are validated with functional experiments using human ESC/iPS and pig EpiSC/iPS. We anticipate that this work will help us develop efficient methods for generating gametes in the laboratory, which will be useful for understanding the basis of infertility, but also with important applications in animal biotechnology.
Given that many aspects of early pig embryo development are similar to human embryos, we use these embryos to uncover fundamental mechanisms of germ line development with relevance to human. The findings from our in vivo characterization are validated with functional experiments using human ESC/iPS and pig EpiSC/iPS. We anticipate that this work will help us develop efficient methods for generating gametes in the laboratory, which will be useful for understanding the basis of infertility, but also with important applications in animal biotechnology.
Mechanisms of gastrulation in mammals
During the first three weeks of development the human fetus becomes almost completely formed. Hitherto, most of our understanding of development is inferred from mouse studies, because human embryos cannot be studied after they implant around of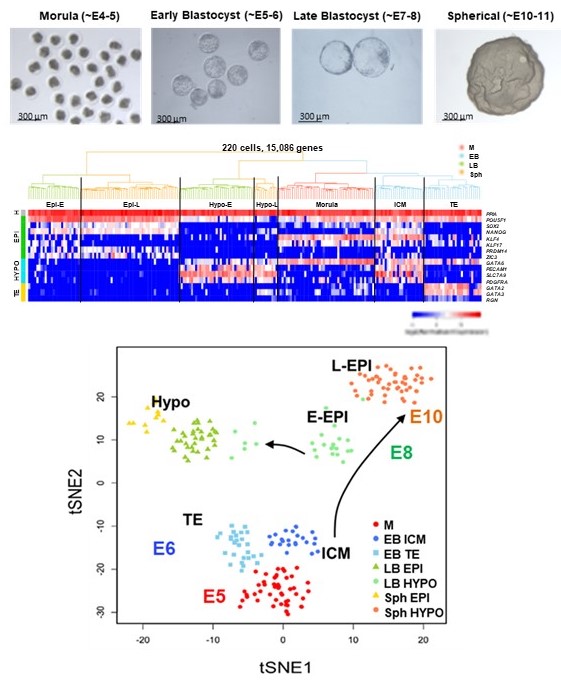 day 9 after fertilization. The pig has recently emerged as a valuable model species that can inform on fundamental mechanisms of human development, is easily accessible and is important in biomedical research. Given that fundamental questions about cell lineage decisions during early gastrulation, also known as the black box of development, remain unanswered, and are critically important for understanding the basis of early embryo mortality and birth defects, we have focussed our efforts in gaining new understanding of pig developmental biology. Using state of the art transcriptomics we demonstrated that pig embryos can be used to broaden the knowledge of human development (Ramos-Ibeas et al., Nature Communications 10:500, 2019), and represent an accessible and ethically acceptable source of embryos for research. Furthermore, given that the pig is an excellent model for biomedical research, increasing our knowledge of its developmental biology will have an impact in human regenerative medicine and our understanding of human genetic diseases.
day 9 after fertilization. The pig has recently emerged as a valuable model species that can inform on fundamental mechanisms of human development, is easily accessible and is important in biomedical research. Given that fundamental questions about cell lineage decisions during early gastrulation, also known as the black box of development, remain unanswered, and are critically important for understanding the basis of early embryo mortality and birth defects, we have focussed our efforts in gaining new understanding of pig developmental biology. Using state of the art transcriptomics we demonstrated that pig embryos can be used to broaden the knowledge of human development (Ramos-Ibeas et al., Nature Communications 10:500, 2019), and represent an accessible and ethically acceptable source of embryos for research. Furthermore, given that the pig is an excellent model for biomedical research, increasing our knowledge of its developmental biology will have an impact in human regenerative medicine and our understanding of human genetic diseases.
A link to our single cell Atlas of pig gastrulation (Simpson et al., 2024, Nat Comms,15:5210) can be found here: https://www.shinyapps.io/
Stem Cell Technologies in livestock species
We have gained detailed understanding of the developmental trajectories of cells that constitute de pre-implantation pig embryo using single cell genomics (Ramos-Ibeas et al., Nature Communications 10:500, 2019). With this knowledge we have developed novel methods for derivation of embryonic stem cells from diverse domestic animal embryos (Kinoshita et al., Development , 148 (23): dev199901, 2021). These cells can be established easily and can differentiate into all cell lineages upon specific cues.
We use these cells to study gastrulation in vitro via generation of gastruloids, an alternative model for functional investigation of gene function during development. This new resource enables high-throughput investigations into fundamental questions of embryo development, and can inform on cell fate specification programs in different animals.
Cultivated meat from livestock stem cells
The new cell lines generated in our laboratory can robustly differentiate into specific line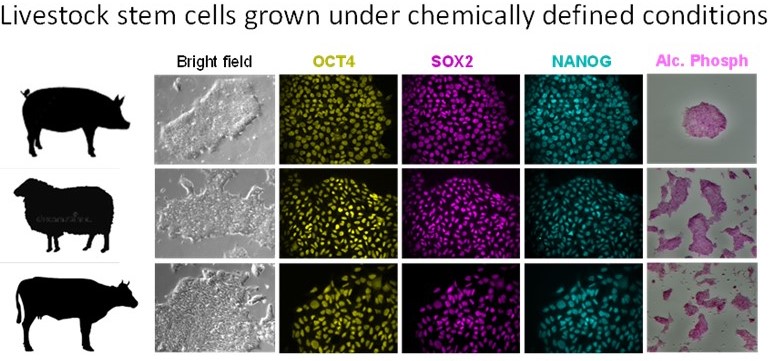 ages. We are currently developing technologies to generate muscle and adipose progenitors to create cultivated meat. We aim to develop this source of nutritious animal proteins as a complement to conventional meat. We expect that this new form of food production will help alleviate the increasing demand for meat worldwide, contributing to a more sustainable production (by reducing the needs of additional land and water). This early stage technology is benefiting of the enormous progress in our understanding of cellular physiology and of the technological advances in engineering and manufacturing.
ages. We are currently developing technologies to generate muscle and adipose progenitors to create cultivated meat. We aim to develop this source of nutritious animal proteins as a complement to conventional meat. We expect that this new form of food production will help alleviate the increasing demand for meat worldwide, contributing to a more sustainable production (by reducing the needs of additional land and water). This early stage technology is benefiting of the enormous progress in our understanding of cellular physiology and of the technological advances in engineering and manufacturing.
Our cells have been used by multiple cultivated meat manufacturing companies worldwide. For details check: https://pluricells.co.uk/
Funding
BBSRC, Innovate UK, Wellcome Trust, Bill & Melinda Gates Foundation.
Current Lab Members:
Doris Klisch (Lab Manager)
Nathan Walters (Technical Specialist)
Dr. Daniel Goszczynski (Research Fellow)
Dr. Luke Simpson (Research Fellow)
Dr. Chuen Li (Research Fellow)
Dr. Noor Shamki (Research Fellow)
Ahmed Ramzy (PhD student)
Zachary Muda (PhD student)
Charlotte Knight (PhD student)
Imge Duru Onol (PhD student)
Katherine Farmery (MRes student)
Former Members:
Sebastian Lazar, MPhil (2008)
Xiaoni Sun, MRes (2010)
Alejandro Contreras, PhD (2011), Senior scientist at University of Mexico
Somsin Petyim, PhD (2013), Head of Obs & Gynaecology, Bangkok, Thailand.
Griselda Valdez Magana, PhD (2014), Senior scientist at University of Mexico
Sergio German, PhD (2015)
Dr. Adam Hirst (Postdoc) Senior Scientist at Stem Cell Technologies.
Haixin Zhang, PhD (2016), postdoc at University of Cambridge.
Sidar Bereketoglu, PhD (2016), group Leader at university of Ankara, Turkey.
Choulia Mola, PhD (2017)
Fabio Amaral, PhD (2017)
Darren Crowley, PhD (2018), senior scientist at Ely Lilly, Ireland.
Haitham Alhilfi, PhD (2018), Academic in Irak.
Judith Gomez-Martinez, PhD (2018), human embryologist at IVF clinic.
Liam Wood, MRes (2018)
Dr. Priscila Ramos-Ibeas (Postdoc), group Leader at INIA Madrid, Spain
Dr. Sarah Withey (Postdoc), Research Fellow, at University of Brisbane, Australia.
Qifan Zhu, PhD (2021), Postdoc at CEMC, CAS, Shanghai, China.
Sophie Kraunsoe (PhD student at the Crick Institute)
Dr. Andrew Strange (Bioinformatics core, University of Sheffield)
Future Research
./.