Genotyping Resources
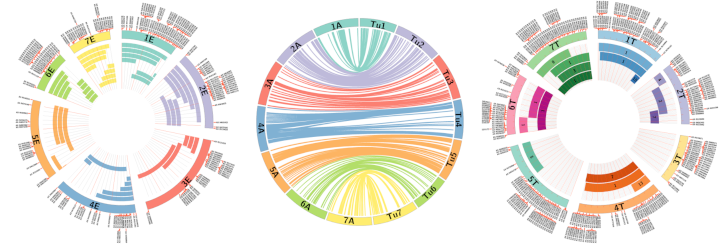
Axiom® Wheat Relatives Array data
This 36K array has been used for genotyping of all back-cross populations generated so far between the following wild relative species and hexaploid wheat:
Details of array content, including probes, sequences and map positions on the wheat genome assembly Refseq v1.0 can be downloaded here  . The array was developed in collaboration with the University of Bristol. More information about the SNPs on the array and development of other molecular marker resources in wheat can be found on the CerealsDB website.
. The array was developed in collaboration with the University of Bristol. More information about the SNPs on the array and development of other molecular marker resources in wheat can be found on the CerealsDB website.
KASP™ Assay Development
A number of chromosome-specific KASP™ assays have been developed for rapid detection of wild relative introgressions into wheat. These assays also allow the identification of the site of introgression in wheat and enable us to differentiate between homozygous and heterozygous introgressions.
Following the publication of this work, the details of all the KASP assays are available below:
Data S1  Genotypes of all the parental and control lines with working KASP assays.
Genotypes of all the parental and control lines with working KASP assays.
Data S2  Details of wild relative species validated for each polymorphic KASP assay.
Details of wild relative species validated for each polymorphic KASP assay.
Data S3  BLASTN and BLASTX results for all chromosome-specific SNPs.
BLASTN and BLASTX results for all chromosome-specific SNPs.
Data S4  BLASTN results for all chromosome-nonspecific SNPs.
BLASTN results for all chromosome-nonspecific SNPs.
Data S5  KASP assay primer details for all selected SNPs.
KASP assay primer details for all selected SNPs.
We are continuously developing new KASP markers to fill in gaps for various wild relatives. For more information on these markers please contact us.