Super Resolution concepts
SIM
SIM Principle
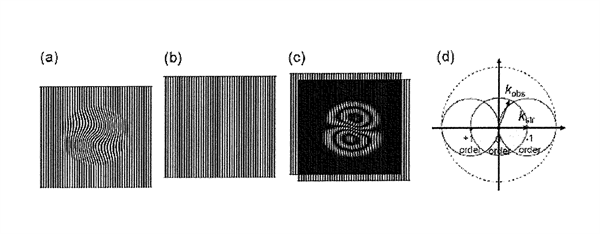
Structured illumination is a wide field technique in which a grid pattern is generated through interference of diffraction orders and superimposed on the specimen while capturing images. The grid pattern is shifted or rotated in steps between the capture of each image set. In the example illustrated below, the image set is composed of 5 individual subsets, each captured after rotating the grid by 60 degrees. Following processing with a specialized algorithm, high-frequency information can be extracted from the raw data to produce a reconstructed image having a lateral resolution approximately twice that of diffraction-limited instruments and an axial resolution ranging between 150 and 300 nanometers. See Zeiss on line tutorial:
Visit Zeiss website
PALM and STORM
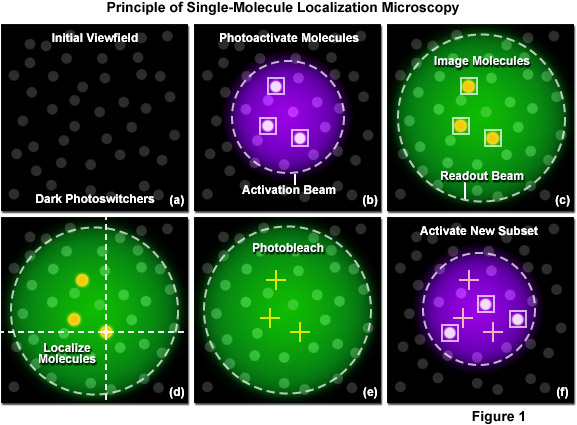
A unique strategy for overcoming the diffraction barrier employs photoswitchable fluorescent probes to resolve spatial differences in dense populations of molecules with super resolution. This approach relies on the stochastic activation of fluorescence to intermittently photoswitch individual photoactivatable molecules to a bright state, which are then imaged and photobleached. Thus, very closely spaced molecules that reside in the same diffraction-limited volume are temporally separated. Merging all of the single-molecule positions obtained by repeated cycles of photoactivation followed by imaging and bleaching produces the final super resolution image. Techniques based on this strategy are often referred to as probe-based super resolution, and were independently developed by three groups in 2006 and given the names photoactivated localization microscopy (PALM), fluorescence photoactivation localization microscopy (FPALM), and stochastic optical reconstruction microscopy (STORM). All three methods are based on the same principles, but were originally published using different photoswitchable probes.
Visit Zeiss website
Photoactivated localization microscopy (PALM) is a super resolution technique that dramatically improves the spatial resolution of the optical microscope by at least an order of magnitude (featuring 10 to 20 nanometer resolution), which enables the investigation of biological processes at close to the molecular scale. The technique relies on the controlled activation and sampling of sparse subsets of photoconvertable fluorescent molecules, either synthetic or genetically-encoded. This interactive tutorial explores the sequential steps involved in creating a PALM image.
View the Zeiss online tutorial
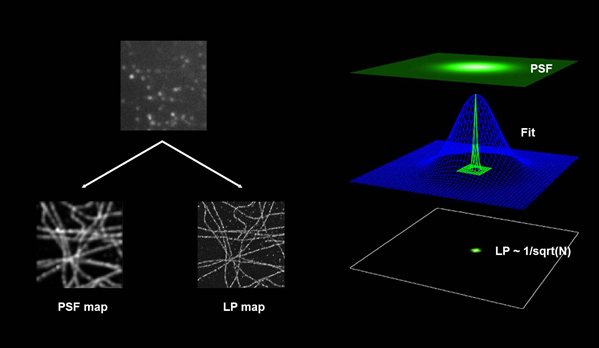
PALM and STORM aim to detect a single fluorescent molecule per point spread function (PSF) at a time. The center of mass of the PSF is calculated from the recorded image, and its position is stored. This process permits a precise localization of the fluorescent signal down to 20 nm. (Carl Zeiss Microscopy GmbH, Dr. Sylvia Münter)
At the end of the experiment the mapped data points / events are displayed into a new image, by showing one pixel per event. Further processing and filtering is applied to correct for oversampling.